Note that unsynchronized pixels (blue) have low coherence throughout the frequency spectrum (left), while synchronized pixels (red) have a high coherence at the ciliary beating frequency (right). Reference pixel used for panel D is shown with a black cross ( D) Schematic depicting how the peak coherence measures ciliary synchronization. ( C) Frequency map of nose pit depicting peak frequency for each pixel. Example pixels used for panel D are shown with crosses. ( B) Raw image frame of a representative light transmission recording in the left nose of a 4-day-old zebrafish larva overlaid with region representing cilia beating (white line). The Fourier transform of pixel intensity time series (top), with peak frequency indicated (bottom). 
As cilia move through a pixel (black rectangle), the pixel intensity fluctuates. ( A) Schematic spectral analysis of a reference pixel. In the scatterplots, the individual data points are light-gray, an average number per fish is in black, and a total average is in red. ( E) To quantify ciliary length, motile cilia expressing GFP ( hspGGFF19B:UAS:GFP) were manually traced from light-sheet microscopy still images. Right: example of a manual tracing of apical surfaces ( D’) the apical surface spans 17.4 µm 2 (☖.31 SD n=11). ( D) To quantify the apical surface of a multiciliated cells, ( foxj1a:GCaMP6s) larvae expressing GFP in multiciliated cells were immunostained for cilia (glutamylated tubulin, magenta), nuclei (DAPI, blue), and cell borders (beta-catenin, red). Insets display for one representative cell the raw fluorescence (top), the signal after applying a bandpass filter (middle), and the detected peaks corresponding to each individual basal body (bottom) ( C’) a cell bears 47.7 cilia (☙.94 SD n=4). ( C) To quantify the number of cilia per cell, basal feet were immunostained with gamma-tubulin (red). ( B’) There is a total number of 50.8 cells per fish (☖.24 SD n=15). Motile cilia are in magenta (glutamylated tubulin), nuclei in blue (DAPI), and multiciliated cells in green ( foxj1a:GCaMP6s).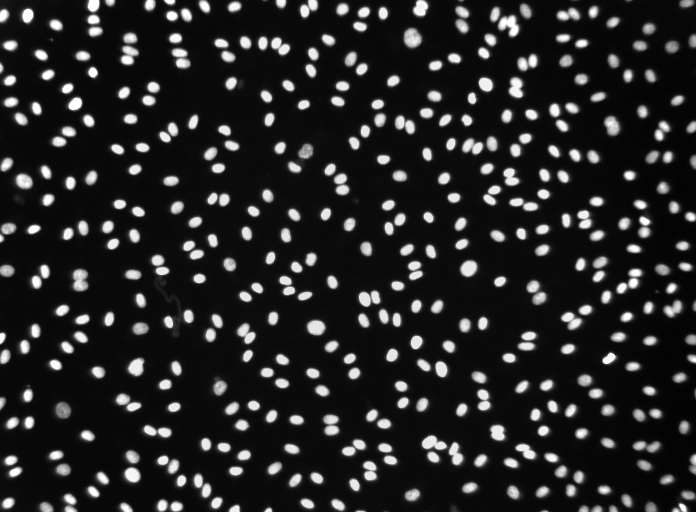 
Ciliated cells in the right nose are indicated in green.( B) A representative example of a left and right 4 dpf zebrafish nose. The approximate location of multiciliated cells and overall fluid flow direction is overlaid on the surface rendering. ( A) Top, front and side view of the surface rendering of a zebrafish head at 4dpf (using a transgenic lines expressing Cherry in all cells, ubi:zebrabow). SD = standard deviation, A: anterior, P: posterior. Shown are the zebrafish nose, clawed frog skin ( Klos Dehring et al., 2013 Kulkarni et al., 2021), mouse brain ventricles ( Redmond et al., 2019), lungs ( Nanjundappa et al., 2019), and oviduct ( Shi et al., 2014). ( E) A graph depicting ciliary density per cell across animals and organs. On average, each cell has 47.7 cilia (☙.9 SD n=4), the apical surface spans 17.4 µm 2 (☖.3 SD n=11), and cilia are 8.83 µm long (☐.86 SD n=38 Figure 1-figure supplement 1B-E'). ( D) A representative example (left) and schematic (right) of a multiciliated cell labelled in the transgenic line hspGGFF19B:UAS:GFP. 
( C) A contour plot showing the average multiciliated cell density (maximum projection) with a total number of 50.8 multiciliated cells per fish (☖.2 SD n=15). DAPI signals highlight the presence of other cell types. Note the lack of multiciliated cells in the center of the nose. In the maximum projection, motile cilia are labeled in magenta (glutamylated tubulin), nuclei in blue (DAPI), and multiciliated cells in green ( foxj1a:GCaMP6s). ( B) A representative example of a left nose marked by a red box in ( A). ( A) Surface rendering of a 4-day-old zebrafish larva (top) and a zoom-in of the nasal cavity (bottom). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
